|
Then they used various methods to identify a subset of conserved elements that accumulated human-specific changes. These analyses started with regions conserved across non-human mammals in order to enrich for functional elements and to increase power to detect acceleration (Box 1). We and the other authors of these studies had a common aim: to identify regulatory elements with human-specific activity (Fig. A number of studies applied different tests to identify HARs either genome-wide or with protein-coding sequences specifically removed from the analysis. Human accelerated regions (HARs) are short, evolutionarily conserved DNA sequences that have acquired significantly more DNA substitutions than expected in the human lineage since divergence from chimpanzees.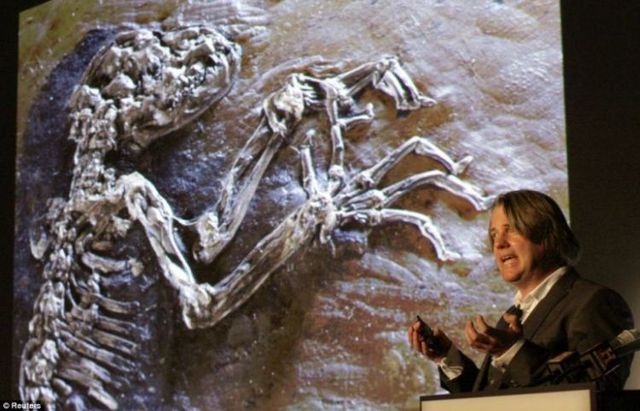 
Hence, human evolutionary genetics has mostly focused on genome regions with many human-specific differences (reviewed in ). Single nucleotide changes can have functional consequences, but currently these are difficult to predict in non-coding regions where small mutations are frequently tolerated and the function of a particular nucleotide is rarely known. 
In this review, we discuss advances to address these barriers with an emphasis on linking sequence to function, complementing other recent papers that explore genetic and regulatory changes in human evolution. Finally, because gene regulation has diverged significantly between primates and model organisms such as mice, zebrafish or flies, it is hard to test hypotheses about the functional effects of regulatory mutations. Furthermore, most uniquely human traits are complex, and there is no doubt that they are encoded by a combination of mutations in different genomic loci. Hence it is difficult to predict the molecular, cellular, and organismal consequences of human-specific regulatory mutations. Second, we know much less about how sequence determines function of regulatory elements compared to protein or RNA genes. The neutral theory of molecular evolution, coupled with redundancy in biological networks, suggests that many human-specific DNA changes had little effect on our biology. First, the non-coding genome is vast, requiring methods to prioritize the mutations that matter. 
The challenge in the post-genomic era has been to determine which of the millions of human-specific non-coding sequence differences are responsible for the unique aspects of our biology. 
Thus, regulatory sequences have great potential to be drivers of human evolution. Furthermore, genes frequently function in many different contexts, and this pleiotropy constrains their evolution compared to regulatory elements, which tend to be more modular. There are many more DNA bases in regulatory regions than in protein-coding genes, making them a larger target for evolutionary innovation. In hindsight, the importance of gene regulation in human evolution is logical. We now know that the vast majority of all genomic changes that happened since the human–chimpanzee ancestor are in non-coding regions, consistent with King and Wilson’s hypothesis that regulatory changes drove the differences between our species. Sequencing our closest living relative, the chimpanzee, begged the question “which genes are different?” Here the answer was predicted a century before and supported by King and Wilson’s 1975 discovery that certain blood proteins have very few amino acid differences between human and chimpanzee. When the human genome was first sequenced, the big question was “how many genes do we have?” Most people guessed too high.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
